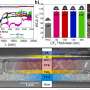
Scientists have successfully sequenced ancient RNA from a juvenile mammoth named Yuka, who lived and died approximately 40,000 years ago in what is now northeastern Siberia. This remarkable discovery sheds light on the last moments of the mammoth’s life, revealing details about its biological processes at the time of death. The RNA was extracted from well-preserved mummified leg tissue, which had remained intact in permafrost for millennia, making it the oldest RNA ever sequenced.
The research team, led by Love Dalén, a professor of evolutionary genomics at the Centre for Palaeogenetics at Stockholm University, published their findings in the scientific journal Cell. They focused on identifying the genes that were active in Yuka’s cells just before it died. “All the cells in an organism have the same DNA,” Dalén explained. “What makes these cells different is essentially the RNA,” which plays a crucial role in determining gene expression in various cell types.
Unlike DNA, which can survive for over a million years, RNA was once thought to be too fragile for long-term preservation. This research, however, demonstrates that under exceptional conditions, RNA can provide valuable insights into ancient life. The team analyzed ten samples of frozen mammoth tissue, including muscle and skin, with only three yielding fragments of RNA. Ultimately, one sample from Yuka produced comprehensive sequencing data that revealed how the mammoth’s genes functioned at the time of its demise.
The data indicated that Yuka was likely close to death, with a noticeable shift in its muscle metabolism. According to Emilio Mármol Sánchez, the study’s lead author from the University of Copenhagen, the findings show the predominance of slow-twitch muscle fibers, suggesting that the mammoth’s muscles were still active in its final moments. Notably, the team identified proteins such as titin, which is related to muscle elasticity, and nebulin, which is involved in muscle contraction.
The significance of these findings extends beyond mere curiosity. Marc Friedländer, a co-author and associate professor at Stockholm University, emphasized that the microRNAs detected in Yuka’s tissues provide direct evidence of gene regulation occurring in real time during ancient epochs. “It is the first time something like this has been achieved,” Friedländer noted.
The implications of this research could potentially revolutionize our understanding of ancient organisms. Erez Lieberman Aiden, a biochemist from the University of Texas Medical Branch, remarked that the ability to detect tissue-specific gene expression is a significant advancement. While the study marks a pivotal moment in the field, Aiden cautioned that it remains uncertain whether RNA will become as vital a reservoir of information as DNA has proven to be for extinct species.
Dalén expressed optimism regarding the future of RNA research, suggesting that the techniques developed in this study could pave the way for new scientific inquiries into the past. He indicated that improvements in methodology are likely, allowing for broader applications in studying ancient viruses and pathogens. The research could also support the goals of Colossal Biosciences, a biotech firm focused on “resurrecting” extinct species like the mammoth by editing the genomes of their closest living relatives.
While Yuka’s RNA represents the oldest recovered to date, it is not the first instance of ancient RNA being sequenced. Previous studies have successfully sequenced RNA from other preserved specimens, including a 130-year-old Tasmanian tiger and a 14,300-year-old wolf. The ability to analyze RNA from ancient tissues opens new avenues for understanding not only extinct species but also the evolutionary history of pathogens.
The groundbreaking nature of this research illustrates the potential for RNA to enhance our comprehension of life in ancient ecosystems and how these findings can inform contemporary scientific endeavors. As methodologies evolve, the insights gained from ancient RNA could lead to significant advancements in genetics, paleogenomics, and even conservation efforts for endangered species.